‘Auxiliary’ Virus Protein Preps Food for Soil-Dwelling Bacteria
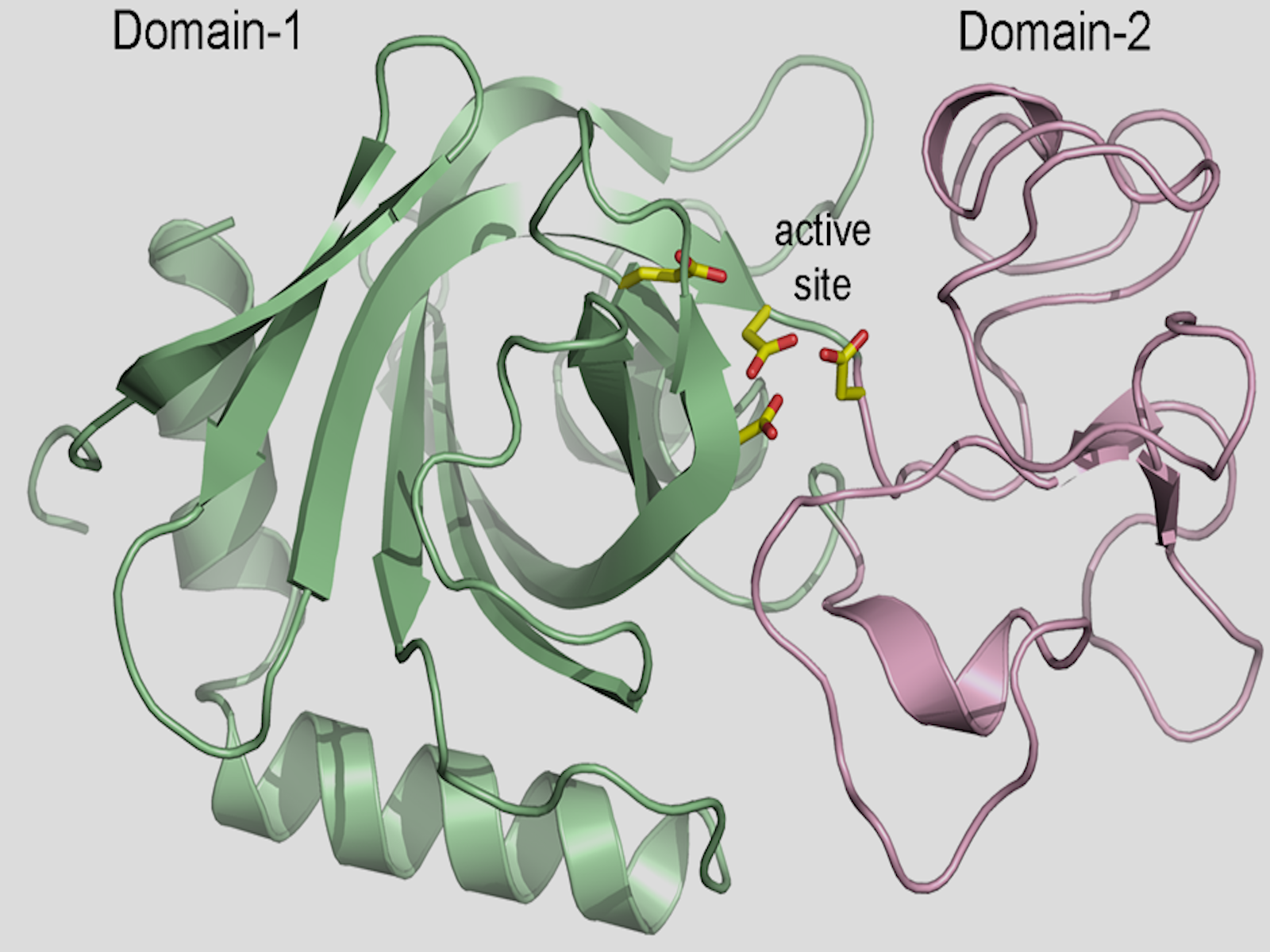
The 3-D structure of a chitosanase–an enzyme that breaks down chitin, a major food source for bacteria in the soil.
(Image by Clyde Smith | SLAC National Accelerator Laboratory)
Scientists have gotten their first glimpse of the structure of a protein whose coding is nestled within a long-hidden landscape: the viruses that inhabit soil and likely play a role in our planet’s carbon cycle.
The protein is the product of what scientists call an auxiliary metabolic gene in viruses. Though viruses carry these genes, researchers know that such “auxiliary” proteins aren’t needed by viruses to replicate. So what are they doing?
A team led by Janet Jansson, a chief scientist at the Department of Energy’s (DOE’s) Pacific Northwest National Laboratory (PNNL), is the first to pin down the structure of such a protein in soil viruses. The team found that the protein is equipped to break down chitin, a common source of food for microorganisms in soil. In other words, the protein prepares a key food source for other microorganisms that are known to affect carbon cycling and other molecular processes in the soil.
The work was published online Sept. 19 in Nature Communications.
Viruses and soil ecology
To solve the protein’s atomic structure, researchers at PNNL searched existing databases for auxiliary genes in soil viruses that scientists had proposed encode enzymes that break down chitin. The gene sequences were then cloned at DOE’s Joint Genome Institute and sent back to PNNL for analysis. The team then crystallized the expressed proteins from the genes using the Stanford Synchrotron Radiation Lightsource at SLAC National Accelerator Laboratory.
Then, the team pieced together more than 5,000 images, each a snapshot of a piece of the protein frozen in time, to reveal the protein’s structure. It’s clear from its structure that the protein is an enzyme known as a chitosanase.
“Our knowledge of viruses and their activity in soil is decades behind our knowledge of other players in soil, such as bacteria,” said Jansson. “This work is the latest clue that viruses may play a very important role in soil ecology.”
In addition to Jansson, other PNNL authors include first author Ruonan Wu, Garry Buchko, Jason McDermott, Kirsten Hofmockel, and John Cort.
The findings were made possible through the collaboration of scientists at several DOE laboratories, including three DOE Office of Science user facilities: the Environmental Molecular Sciences Laboratory, the Joint Genome Institute, and the Stanford Synchrotron Radiation Lightsource. The work was funded by the DOE Office of Science.
To learn more about the findings, click here.
Published: September 20, 2022