Global Science to Unravel Patterns in River Metabolomes
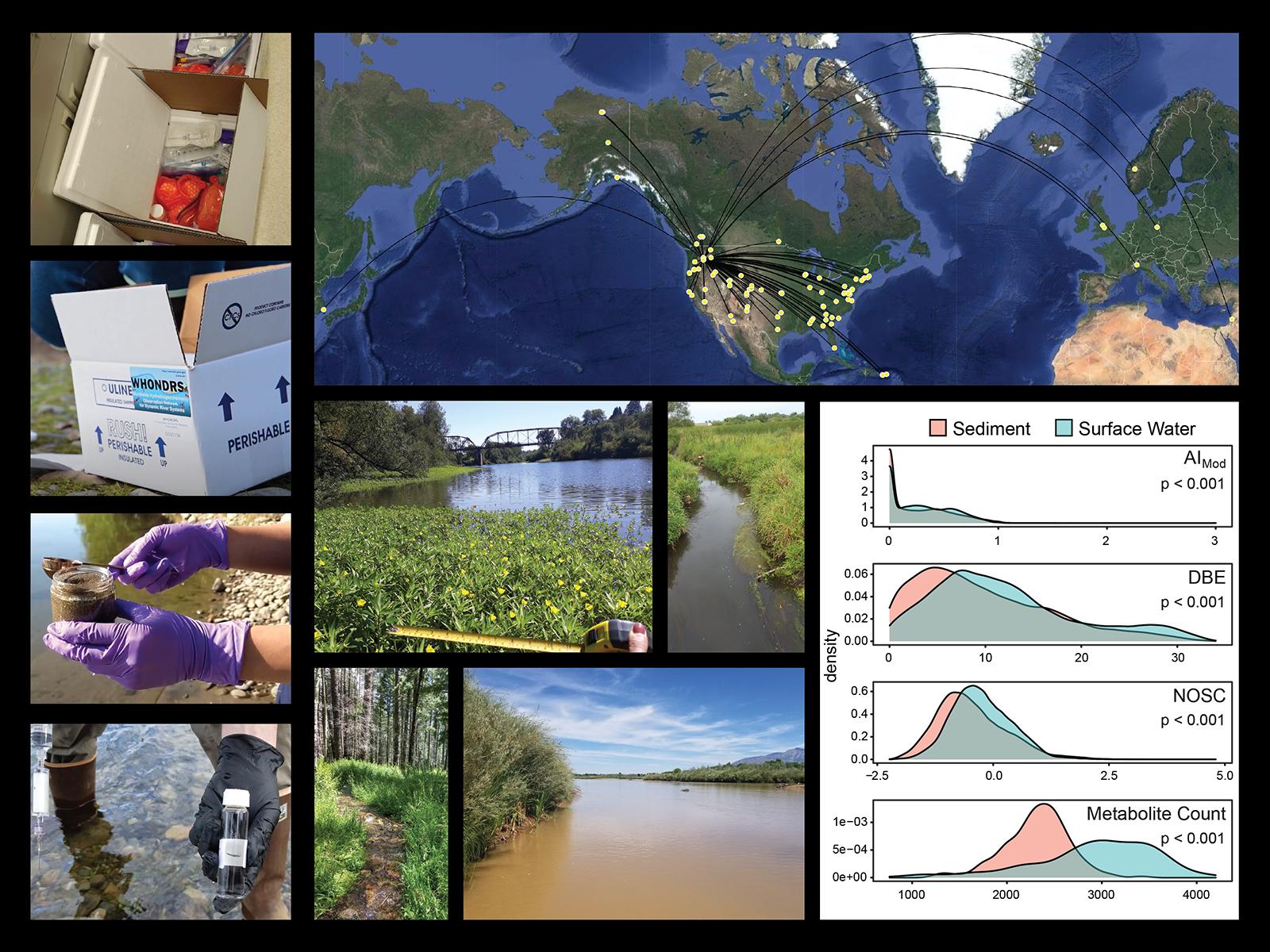
A global effort conducted by the WHONDRS consortium from July to August 2019 used advanced mass spectrometry to characterize metabolomes in surface waters and sediments spanning a range of biomes and stream sizes.
(Composite image by Vanessa Garayburu-Caruso | Pacific Northwest National Laboratory)
The Science
The Worldwide Hydrobiogeochemistry Observation Network for Dynamic River Systems (WHONDRS) is a global consortium of researchers based out of the U.S. Department of Energy’s Pacific Northwest National Laboratory. The consortium uses a standardized approach to understand coupled hydrologic, biogeochemical, and microbial functions in river corridors. Now the group reports the first ultrahigh-resolution analysis of global river corridor metabolomes of both surface water and sediment across rivers spanning a wide range of sizes and ecosystem types. The scientists describe the distribution of key chemical attributes of metabolomes, including East-West gradients, in many metabolomic features across the contiguous United States. They also show that surface water metabolomes are more diverse than those in sediment, possibly suggesting a greater diversity of biological processes occurring in surface waters.
The Impact
Organic matter transformations in aquatic ecosystems are a critical source of uncertainty in global biogeochemical cycles. Environmental metabolomics, or the analysis of organic molecules in environmental samples, help characterize the interactions of organisms within their environment. Environmental metabolomics enabled by ultrahigh-resolution mass spectrometry reveals connections between organic matter character, reactivity, and inferred biochemical transformations within and across localized river corridor ecosystems. These processes, however, are not well understood at the continental-to-global scale. This work provides a foundation for understanding global patterns in river corridor biogeochemical cycles. It also demonstrates that research done following ICON-FAIR principles can enable discoveries that are unfeasible with traditional research models.
Summary
In 2019, WHONDRS worked with the global science community to develop and implement a study based on the ICON-FAIR principles. In this approach, research is designed intentionally to be integrated (I) across disciplines, coordinated (C) with consistent protocols, open (O) by generating data that is Findable, Accessible, Interoperable, and Reusable (FAIR), and networked (N) so that the distributed science community is engaged in the execution of the work. Using these principles, the WHONDRS consortium collected surface water and sediment samples from 97 river corridors in 8 countries within a 6-week period across a wide range of environmental factors such as stream order, climate, vegetation, and geomorphological features. Scientists in EMSL, the Environmental Molecular Sciences Laboratory, a U.S. Department of Energy scientific user facility located at Pacific Northwest National Laboratory, then characterized metabolomes in these samples using Fourier-transform ion cyclotron resonance mass spectrometry.
The scientists described central aspects of the metabolomes, including assigned elemental groups, chemical classes, descriptor indices, and inferred biochemical transformations. Using those features, the scientists then described key metabolome characteristics of surface water and sediment. Finally, they explored spatial patterns across East-West gradients of many of these characteristics within the United States and how they varied among surface waters and sediments. Their work provides a benchmark for understanding global patterns in river corridor organic matter chemistry and highlights the benefits of engaging in ICON-FAIR science to increase transferability of knowledge. This publication was invited to be part of a Special Issue in the journal Metabolites on the Metabolome and Fluxomics.
Funding
This research was supported by the U.S. Department of Energy (DOE), Office of Science, Biological and Environmental Research program as part of the River Corridor Hydrobiogeochemistry Scientific Focus Area at Pacific Northwest National Laboratory. A portion of the research was performed at the Environmental Molecular Science Laboratory, a DOE Office of Science user facility.
Published: January 14, 2022
V.A. Garayburu-Caruso, et al. “Using Community Science to Reveal the Global Chemogeography of River Metabolomes.” Metabolites 10, 518 (2020). [DOI: 10.3390/metabo10120518]