Featured Stories
September 18, 2024
Flexible Circuits Made with Silk and Graphene on the Horizon
September 13, 2024
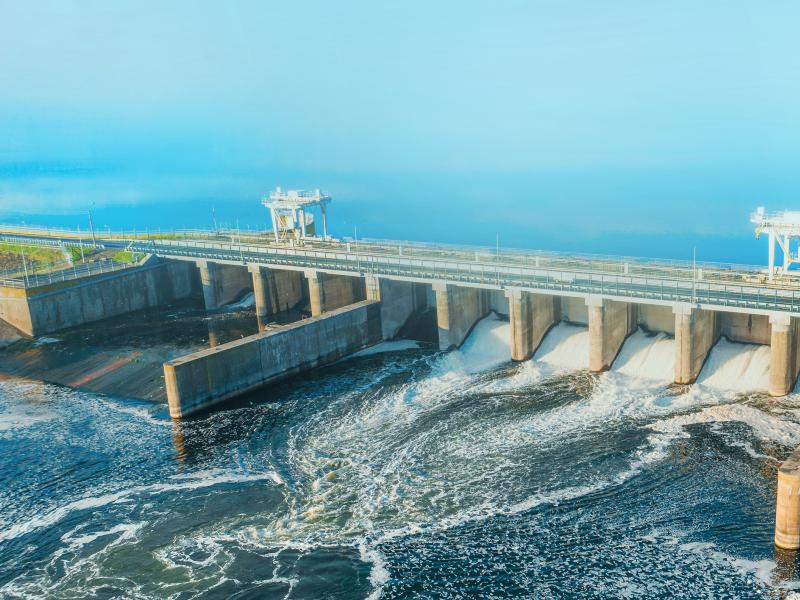
Transmission and Renewables Would Reduce Carbon Emissions, Generation Costs in Western United States
September 10, 2024
PNNL Takes Its Radiation Expertise to Space
September 5, 2024
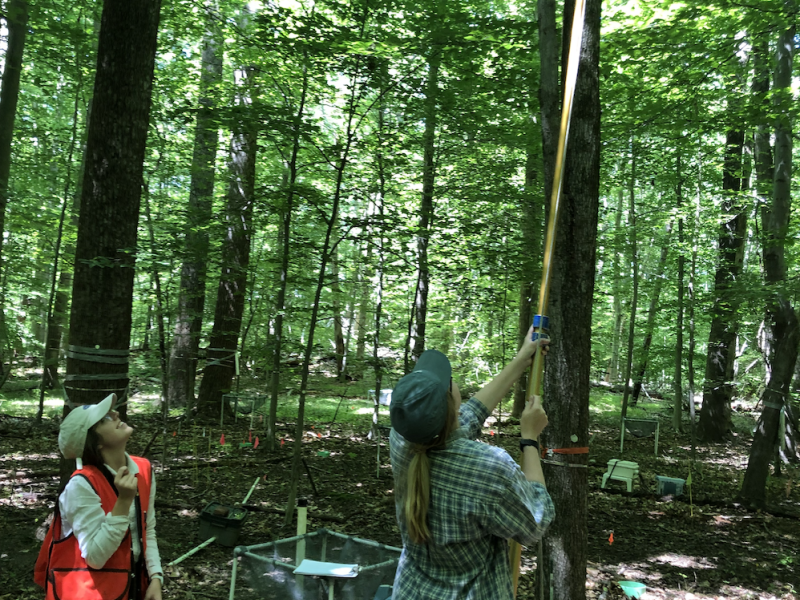
Research Vessel Resilience Charts Course to the Future of Marine Research
Subscribe
to receive PNNL
news by email:
Latest Stories
513 results found
Filters applied: Biology, Hydropower and the Electric Grid