Using the Metaproteome to Uncover the Microbiome of the Complex Soil Environment
Scientists developed a new deep learning computer model that can use information about proteins to more accurately identify the microbial community in a soil sample
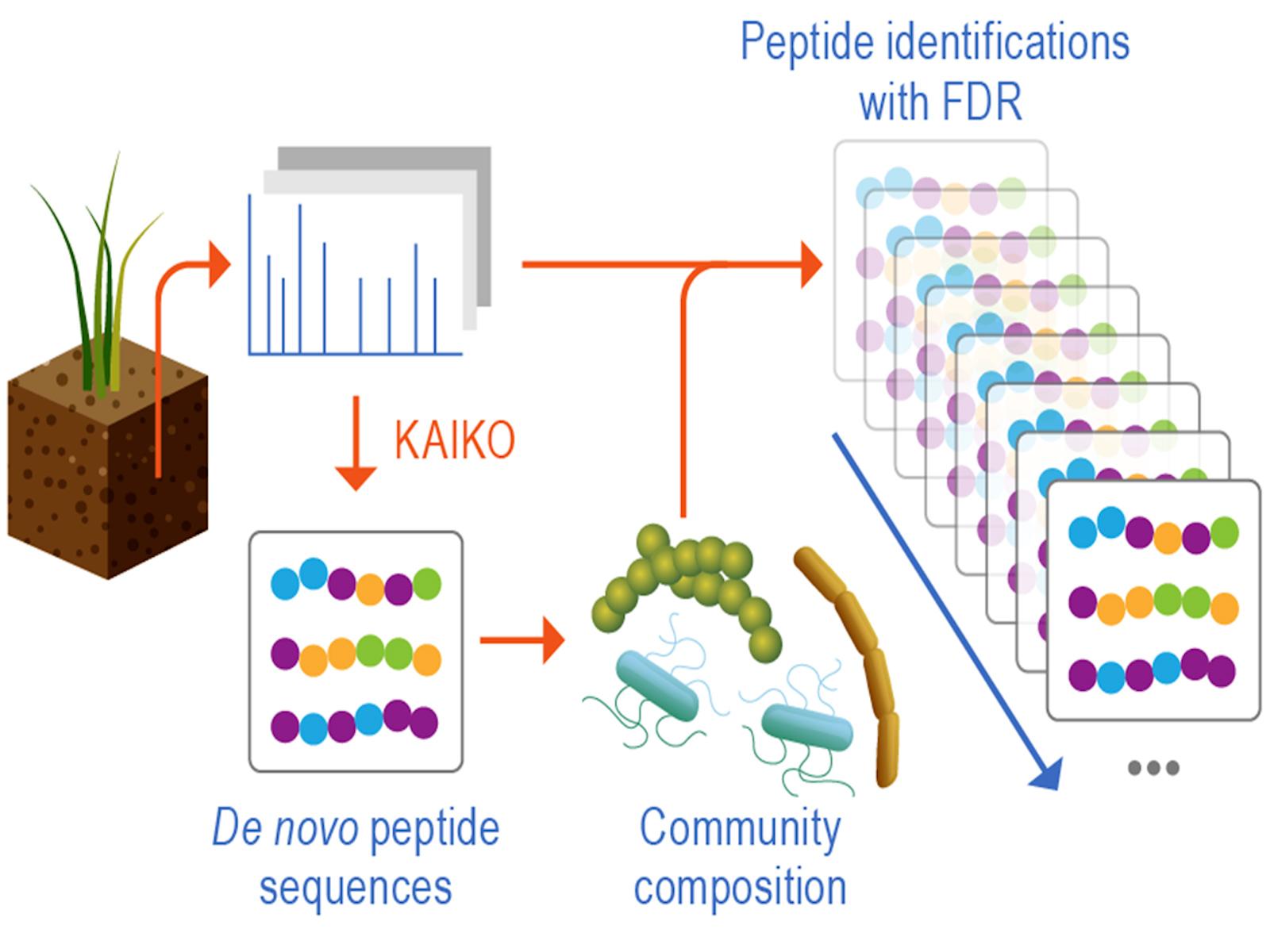
Scientists developed a new approach to analyze environmental samples, including creation of a sample-specific protein sequence database that can identify genes and proteins from microbes more accurately than existing digital tools. Description of text in graphic: The Kaiko deep learning tool workflow include de novo peptide sequences, community compositions, and peptide identifications with false discovery rate (FDR).
(Graphic: Pacific Northwest National Laboratory)
The Science
Recent advances in soil microbiology have allowed scientists to peer deep inside environmental samples to begin to understand the biological functions carried out by microbes in soil. At such a tiny scale, the only way to identify these microbes is to compare their genes and proteins to database information of previously identified species. But what if such databases aren’t readily available, as is often the case in laboratories with limited ability and funding, to access the latest technologies and data systems? Using a new approach, a broad spectrum of scientists can now generate a protein database directly from proteomics data gathered from a specific soil sample. A key element of this new approach is a digital tool called Kaiko, a deep learning computer model that has significantly improved accuracy compared to currently available digital tools for generating a protein database.
The Impact
The soil microbiome carries out many functions important on a global scale, including cycling of carbon and other nutrients as well as supporting plant growth. Although research has greatly expanded the number of microbiome species that can be identified by their genes and proteins, significant practical and financial barriers still prevent many soil scientists from always having access to an assembled and well-annotated database of proteins for comparison. Therefore, tools that can create a database for soil samples as soon as possible are a significant benefit to scientists studying the soil microbial community, and could open the door to a more complete understanding of the role that a particular microbial community plays in climate change and agriculture.
Summary
Like the Japanese deep ocean submersible used to explore the Marianas Trench after which it was named, a newly developed digital tool called Kaiko dives deep into data to identify and learn complex patterns. After creating Kaiko, the new digital tool had to be trained using peptide sequence data from mass spectrometry. To teach Kaiko about proteins, scientists mined the extensive archive of mass spectrometry data obtained and maintained by EMSL, the Environmental and Molecular Sciences Laboratory, a Department of Energy (DOE) Office of Science user facility. The training data included a set of 5 million sample matches from 55 diverse microorganisms across 9 phyla. With the EMSL data as its training set, Kaiko successfully identified organisms directly from the proteomic data from natural and synthetic soil samples. Finally, the team of scientists involved in this research created a process to generate a database of all the proteomic data from a sample, or a metaproteome, using Kaiko, and tested the process on native soils collected from a site in Kansas. The process identified all highly abundant microbes and uncovered several additional species. The new digital tool will allow a greater number and variety of scientists around the world to study the soil microbiome.
Contacts
Kristin Burnum-Johnson, EMSL and Pacific Northwest National Laboratory, Kristin.Burnum-Johnson@pnnl.gov
Samuel Payne, Brigham Young University, sam_payne@byu.edu
Funding
Funding for this project was provided by the DOE Office of Science, Biological and Environmental Research program, Early Career Research Program, and Pacific Northwest National Laboratory’s Deep Learning for Scientific Discovery initiative. Proteomics data used in this manuscript were generated in EMSL, the Environmental Molecular Sciences Laboratory, a DOE Office of Science user facility.
Published: September 5, 2023
J.-Y. Lee, et al., “Uncovering Hidden Members and Functions of the Soil Microbiome Using De Novo Metaproteomics.” Journal of Proteome Research 21, 2023-2035 (2022). [DOI: 10.1021/acsjproteome.2c00334]