Northwest Biopreparedness Research Virtual Environment (NW-BRaVE) Initiative
Northwest Biopreparedness Research Virtual Environment (NW-BRaVE) Initiative
Supporting innovations in data analytics, instrumentation, and experimental techniques for the development of new approaches for disease detection, targeted therapy design, and antipathogen materials.

Research topics
Lab-Level Communications Priority Topics
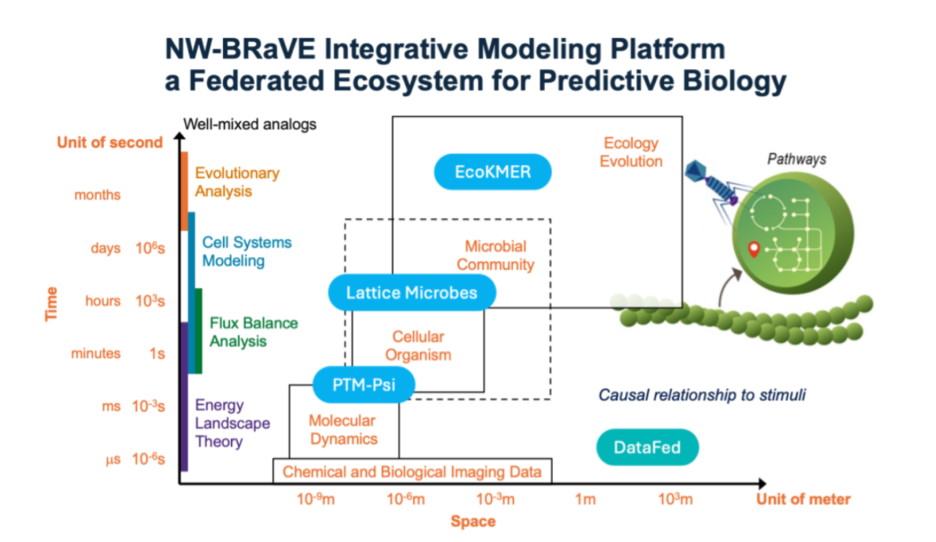
Predictive Phenomics