Featured Stories
December 9, 2025
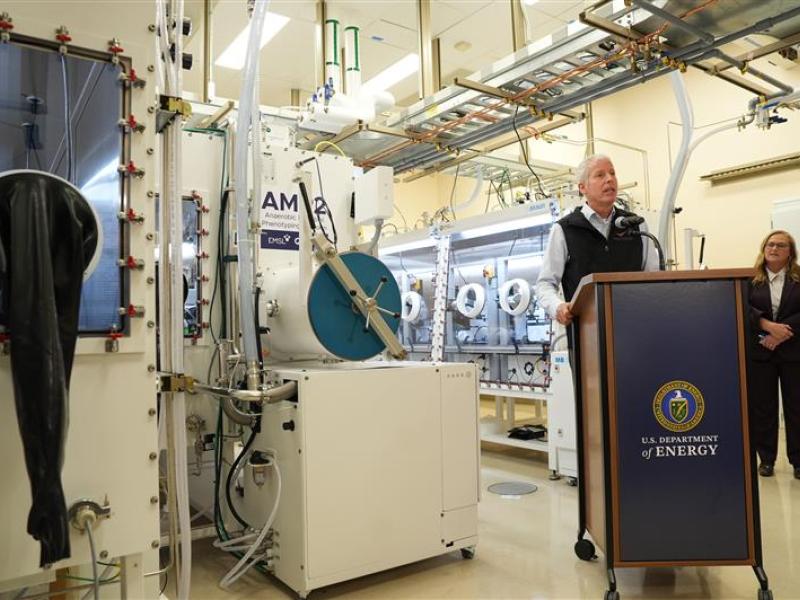
First Testing of Grid-Scale Battery Technology Begins at the Grid Storage Launchpad
December 4, 2025
Department of Energy, PNNL Partner to Power the Nation’s Bioeconomy
December 11, 2025
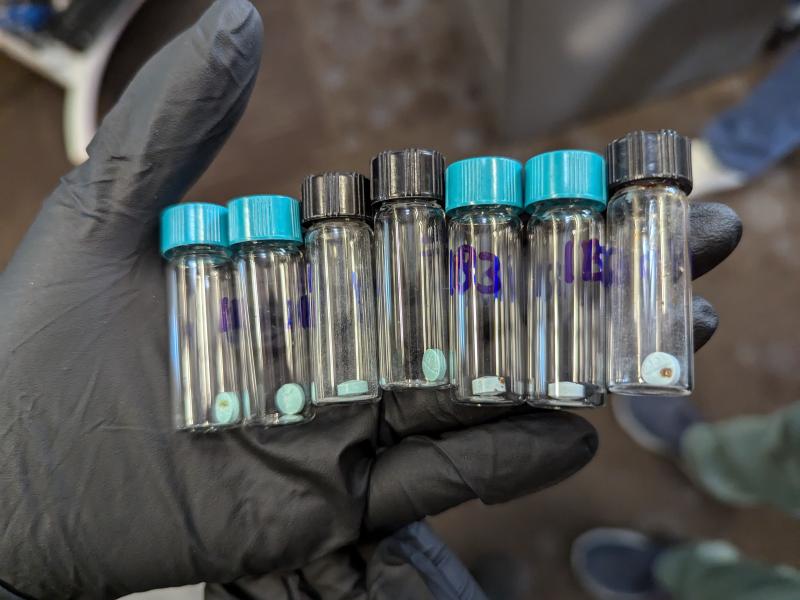
From Molecules to Batteries and Beyond: Celebrating PNNL’s Energy Storage Legacy
November 24, 2025
Energy Department Launches ‘Genesis Mission’ to Transform American Science and Innovation Through the AI Computing Revolution
Subscribe
to receive PNNL
news by email:
Latest Stories
172 results found
Filters applied: Subsurface Energy Systems, Integrative Omics, Soil Microbiome