Model Microbial Consortium Simplifies Soil Microbiome Complexity
A small collection of naturally interacting soil microbes could enable new studies of community interactions and metabolism
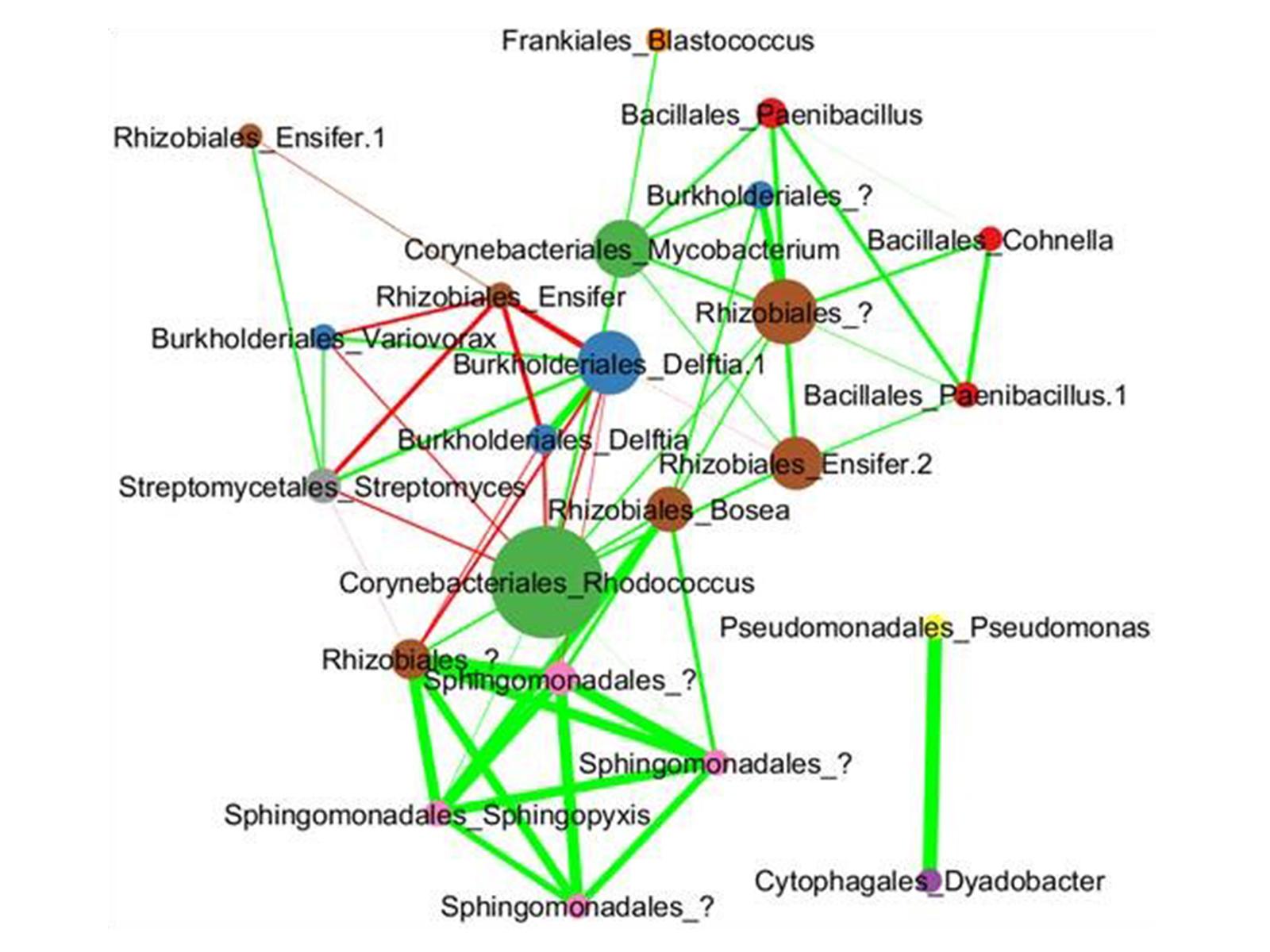
Each circle represents a microbial species within the model consortium MSC-1. Lines between species indicate putative interaction, with the thickness of the line indicating the strength of the interaction. The size of the circle indicates the species’ predicted importance.
(Image courtesy of Ryan McClure | PNNL)
The Science
The soil microbiome plays a major role in the cycling of carbon and nitrogen, and also in promoting plant health. However, the high diversity of the soil microbiome and the complexity of the soil habitat make it extremely difficult to understand the myriad of species interactions involved in organic carbon decomposition. Researchers at the U.S. Department of Energy’s Pacific Northwest National Laboratory reduced this complexity by developing a model soil consortium that is simpler, yet still representative of the native soil. This consortium, called MSC-1, was naturally enriched to contain dozens of interacting species. This collection of several interacting species allows for more in-depth analysis of specific interactions and metabolic exchanges that simply would not be possible when applied to a native soil microbiome.
The Impact
This unique model soil community, with a tractable number of species, will serve as a valuable resource for the larger science community to understand how soil microorganisms interact to carry out important processes in the native soil environment. MSC-1 was able to be stored and revived while retaining the same microbial composition. This feature enables the consortia to be used as tools for future use or shared with other scientists.
Summary
Researchers developed a model soil microbial consortium that is simpler and more amenable to experimental analysis compared to the native soil microbiome. They collected samples from an arid grassland soil and enriched the microbial communities by growing them on chitin as the main carbon source. Chitin is a complex organic molecule common in soil that often requires metabolic coordination from several microbial species for decomposition.
This enrichment approach resulted in defined consortia with a relatively low number of species (30-50) and large phylogenetic diversity. The resulting consortia could be stored and revived while retaining their community compositions, greatly increasing their use as tools for the research community at large. One of the consortia that was particularly stable was chosen as a model soil consortium (MSC-1) for further analysis.
The researchers studied species interactions within MSC-1 using both pairwise co-cultivation in liquid media and during growth in soil under several stresses found in native soil. Co-abundance analyses highlighted interspecies interactions and helped to define keystone species, including Mycobacterium, Rhodococcus, and Rhizobiales taxa. These experiments demonstrate the success of naturally enriching a community of interacting species that can be stored, revived, and shared.
Contact
Ryan McClure
Pacific Northwest National Laboratory
ryan.mcclure@pnnl.gov
Funding
This research was supported by the U.S. Department of Energy, Office of Science, Biological and Environmental Research Program and is a contribution of the Scientific Focus Area “Phenotypic response of the soil microbiome to environmental perturbations.”
Related Links
Amplicon sequence data has been deposited at the PNNL DataHUB repository here: https://doi.org/10.25584/WAIsoCMSC1/1635272 and is available for download under the project accession 10.25584/WAIsoCMSC1/1635272. The version described in this article is the first version. Package includes amplicon 16S rRNA fastq files, supplementary information, MIxS_MIMARKS.soil.5.0 metadata information, R markdown, and package “Read Me” file. The R markdown processing scripts used to process the data and build graphs are available at: https://github.com/dtnaylor124/ModelMicrobialCommunity
Published: November 9, 2020
R. McClure, et al., “Development and Analysis of a Stable, Reduced Complexity Model Soil Microbiome.” Frontiers in Microbiology 11, 1987 (2020). [DOI: 10.3389/fmicb.2020.01987]