Integrative Network Modeling Reveals Key Drought-Associated Genes Key in the Soil Microbiome
Molecular and gene networks were combined to better understand how soil microbial communities respond to changes in water levels and nutrient sources
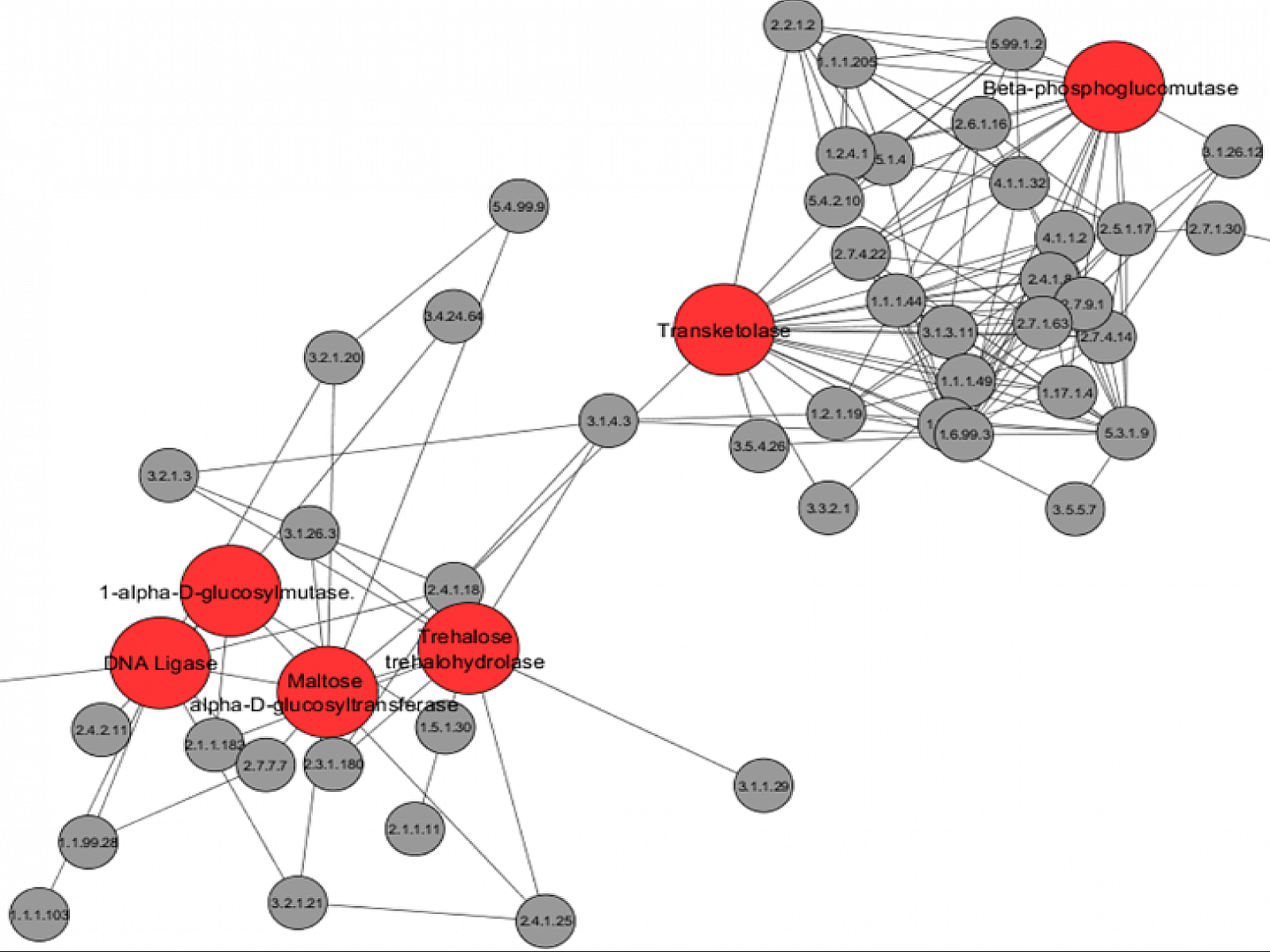
A new network analysis of microbial molecular responses to environmental stresses reveals connections between bacterial genes that respond to drought conditions (large red circles) and other genes and processes (smaller grey circles). Image from R.S. McClure, et al., “Integrated network modeling approach defines key metabolic responses of soil microbiomes to perturbations.” Scientific Reports 10, 10882 (2020). [DOI: 10.1038/s41598-020-67878-7]
The Science
Harnessing the soil microbiome to enhance ecosystem services, like plant productivity or bioenergy production, requires understanding how soil microbiomes respond to environmental stresses, such as flood, drought, or changing nutrient levels. In this study, researchers examined how the soil microbiome responds at genetic and metabolic levels to changes in water content and nutrient sources. This integrated network analysis identified unique sets of genes and metabolic reactions that are expressed only under wet, dry, or high nutrient conditions. When focusing on these unique genes and pathways, the analysis showed that genes associated with dry soil conditions are central to the soil microbiome's response to environmental shifts.
The Impact
The soil microbiome promotes plant health and affects the cycling of carbon through the ecosystem. Here, researchers examined how many different genes and molecules expressed by soil microbes are related to each other and to certain environmental conditions. They were also able to identify how individual genes occupied key positions in a functional network. The researchers found that the soil microbiome particularly responded to dry conditions. This knowledge will help in future efforts to harness the soil microbiome for optimizing plant productivity under drought conditions.
Summary
It is challenging to untangle the complex response of the soil microbial community to environmental change, partly due to the absence of modeling frameworks that can predict how environmental changes in soil can lead to changes in the microbial community’s function and role in promoting soil health. To fill this gap, researchers performed a combined analysis of metabolic and gene co-expression networks to explore how the soil microbiome responds to changes in moisture and nutrient conditions. The integrated modeling approach revealed previously unknown, but critically important, aspects of the soil microbiomes’ response to environmental perturbations, including soil desiccation. Incorporation of metabolomic and transcriptomic data into metabolic reaction networks identified condition-specific signature genes that are uniquely associated with dry, wet, and glycine-amended treatments. A subsequent gene co-expression network analysis revealed that dry-associated genes, in particular, are central to the network; this means they are especially critical to the soil microbial community’s response to changing conditions. These results indicate the occurrence of a system-wide microbiome metabolic coordination when soil microbiomes cope with moisture or nutrient perturbations. Importantly, this approach to analyzing large-scale multi-omics data from a natural soil environment is applicable to other microbiome systems for which genomic and metabolite data are available.
PNNL Contacts
Kirsten Hofmockel, Pacific Northwest National Laboratory, kirsten.hofmockel@pnnl.gov
Funding
This research was supported by the U.S. Department of Energy, Office of Science, Biological and Environmental Research Program, as part of the Genomic Science Program, and is a contribution of the Pacific Northwest National Laboratory Soil Microbiome Scientific Focus Area "Phenotypic Response of the Soil Microbiome to Environmental Perturbations."
Published: July 24, 2020
R.S. McClure, et al., “Integrated network modeling approach defines key metabolic responses of soil microbiomes to perturbations.” Scientific Reports 10, 10882 (2020). [DOI: 10.1038/s41598-020-67878-7]