Multi-omics Data are Key to Advancing Reactive Transport Models
An overview of recent advances in microbial reactive transport models identifies omics data as the ‘current frontier’ for understanding system-scale microbial behavior and dynamics.
The Science
Reactive transport models (RTMs) are used to describe and predict the distribution of chemicals in time and space, in both marine and terrestrial (surface and near-surface) environments where microbially-mediated processes govern biogeochemical patterns. Yet, challenges exist in modeling microbially-driven systems, as well as inintegrating data across the vast range of scales relevant to models of biogeochemical cycling.
In the April 2019 topical issue of the journal Elements on reactive transport modeling, Tim Scheibe of Pacific Northwest National Laboratory (PNNL) and coauthor Chistof Meile of the University of Georgia discuss common approaches that have been used to incorporate microbial community interactions and their influence on geochemical processes in RTMs, and future opportunities to leverage new instrument and data capabilities—including multi-omics—to create new and more realistic modeling approaches.
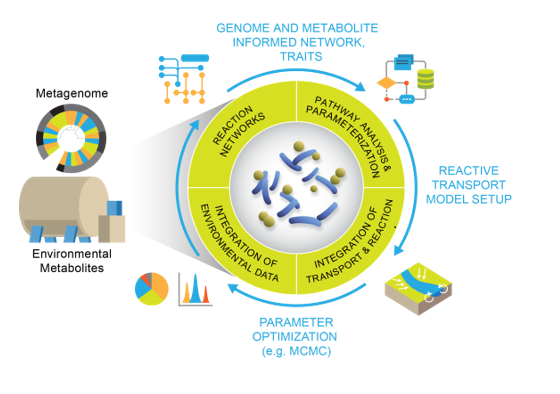
In particular, the authors argue that omics-informed microbial models will help advance understanding of how complex microbial communities respond to environmental changes. These new models will also help identify microbial impacts on local and global elemental cycling, the fate of contaminants, redox transformations, and other processes mediated by microorganisms.
The Impact
Integrating omics data into RTMs will improve predictive understanding of critical watershed processes such as carbon and nitrogen cycling within those watersheds and more broadly. Omics-informed modeling will also reveal how critical microbial processes change in response to environmental perturbations—an urgent imperative for watersheds subject to increasingly frequent or sustained perturbations.
Summary
Representation of microbial processes in RTMs has advanced significantly over the past few decades, accounting for dynamic changes in biomass, functional regulation in response to environmental changes, and thermodynamic constraints. Current RTMs represent microbial functions with greater process fidelity and reduced empiricism.
The authors say that incorporating multi-omics data is a current frontier in RTMs, and offers great potential for improving scientific understanding of microbial processes and predictive modeling. To that end, they are engaged in research to integrate complex metagenomics, metabolomics, and other omics data into reaction network models. In turn, these can be linked with state-of-the-art RTMs in order to simulate system-scale behavior.
In the article, the authors introduce relevant case studies and discuss ways to integrate multi-omics data to inform and validate RTMs. Their results advance and enhance those modeling capabilities by identifying and promoting how to integrate multi-omics data into microbial models.
The result, the authors say, will be an improved predictive understanding of critical watershed processes such as carbon and nitrogen cycling within specific watersheds and more broadly. Omics-informed modeling will also reveal how critical microbial processes change in response to environmental perturbations.
Contact
Timothy D. Scheibe, Pacific Northwest National Laboratory, Tim.Scheibe@pnnl.gov
Funding
U.S. Department of Energy, Office of Science, Biological and Environmental Research (BER), Subsurface Biogeochemical Research (SBR) Program.
Revised: September 6, 2019 | Published: September 9, 2019
Meile, C. and Scheibe, T.D. 2019. Reactive Transport Modeling of Microbial Dynamics, Elements, 15(2): 111-116, doi: 10.2138/gselements.15.2.111