Advanced Computing, Mathematics and Data
Research Highlights
February 2015
Cross-institutional Team Demonstrations Tackle Big Data Challenges in Materials Science
Collaborative environment for experimental facilities, computational modeling, and federated data science capabilities drives innovation
The synthesis and functionality of energy storage and conversion materials is a key research area for the U.S. Department of Energy’s Office of Basic Energy Sciences (DOE-BES). As part of a series of demonstrations showcased at DOE’s booth during the SC14 conference in New Orleans, Kerstin Kleese van Dam, Chief Scientist and Data Services Team Lead at Pacific Northwest National Laboratory, led talks showing how the combination of leading-edge microscopy facilities, computational modeling, and federated data science capabilities—as well as cross-domain collaborations—can significantly advance fundamental scientific understanding and control of the critical materials processes in these systems. The demonstrations at SC14 highlighted the ongoing work of DOE’s Data Science Centers, which are uniting national laboratories, academic institutions, and international partners in an effort to improve methods for collecting, analyzing, and sharing Big Data. PNNL scientists are contributing on several teams, including those representing BES, DOE’s Office of Biological & Environmental Research, and a team focused on overall data infrastructure challenges.
Pushing Frontiers
For the SC14 demonstration, Kleese van Dam described how her data team chose an area where many of the scientific user facilities are pushing the frontiers of science and technology: understanding the nucleation and growth of nanoparticles from solution. This research allows the relationship between nanoparticle size, shape, composition, and functionality; the processes behind synthesis and emergent mesoscale functionality; and the origin of degradation mechanisms in next-generation batteries, all to be investigated.
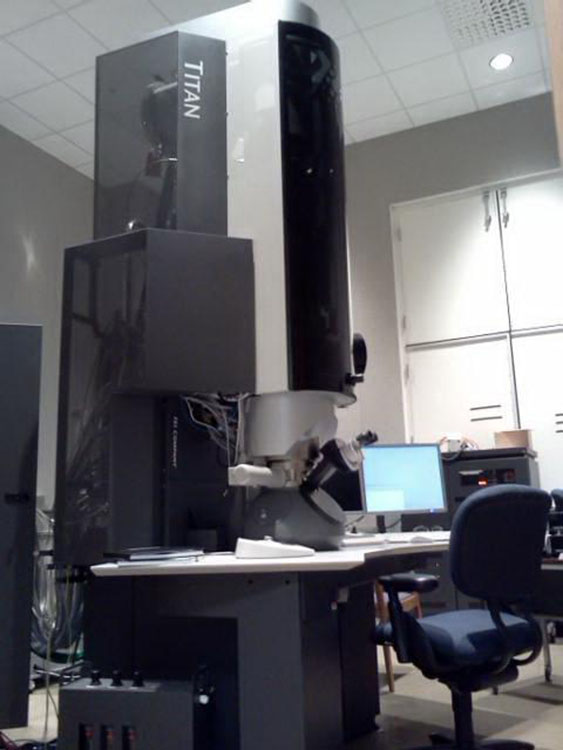
As with many other experimental technologies, electron microscopy is undergoing a technology-driven transformation. Where, in the past, experimental methods did not possess the spatial or temporal resolution to capture physical, chemical, and biological processes in sufficient detail, new experimental in situ and in operando modalities promise to enable scientists to capture evolving processes. Technically, systems are moving from being capable of capturing 100 images per day to microscopes that now capture 100 to 1,000 images per second, with further substantial increases expected from new instruments, including dynamic transmission electron microscopes. Crucially, this development, in combination with in situ analysis and interpretation, may provide the necessary capabilities for experimental steering and, ultimately, control of transformations, leading not only to faster scientific progress and better experimental results, but opening up the possibility of completely new scientific discoveries.
This transition in experimental modalities goes hand in hand with a dramatic increase in data rates and volumes, which render existing analytical tools insufficient. Today, the experimental community is ill prepared to adapt to its fast-changing analysis needs. For the most part, existing tools are developed specifically for individual instruments and are not scalable. High-performance computing (HPC)-enabled analysis and simulation could have a major impact on the design and interpretation of experiments as part of the materials design process. However, their use today is limited by the high entry barriers for non-experts, both in their development and use. Therefore, it is essential to create a community environment where experimentalists, modelers, and data analysts can easily work together to create new scalable solutions and provide everyone with easy access to the resulting tools and data.
A ‘Premier’ Demonstration
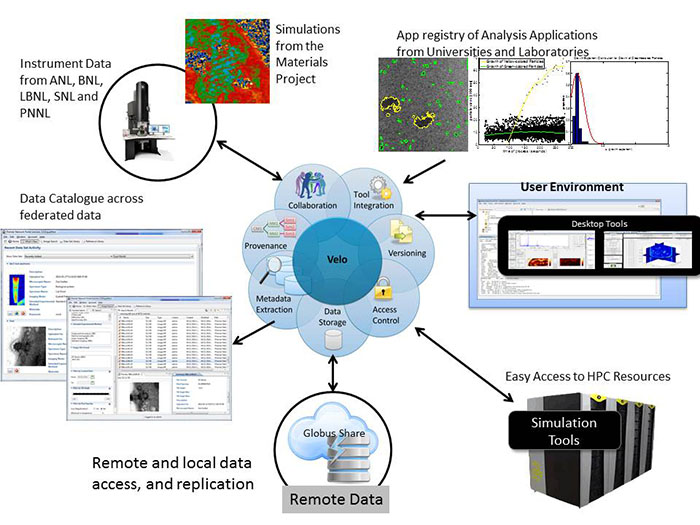
The core demonstration environment was based on the PNNL-developed Premier Portal service, a collaborative environment for microscopists currently supporting 30 institutions organized in the Premier Network. A further important component was the Argonne National Laboratory (ANL)-developed Globus Online data transfer service. The Premier Portal is a customization of Velo, a reusable, scalable, domain-independent infrastructure for managing scientific work that supports the entire scientific project life cycle, including data management, modeling and simulation, visualization and analysis, validation, reporting, archives, publishing, and discovery. Velo is built upon extensible open-source technologies to create a collaborative core platform that can be tailored to specific scientific uses and deployed to new sites within weeks. Velo has been developed over the past 10 years and is in production use by 15 separate projects and services supported by the U.S. Department of Homeland Security, Environmental Protection Agency, and DOE. Globus Online provides fast and reliable data transfer services, making data accessible and sharable across distributed environments.
For this demonstration, the focus was on showcasing the impact of two key new features: a federated data sharing and analysis environment, as well as providing easily usable HPC-enabled data analysis.
Federated Infrastructure
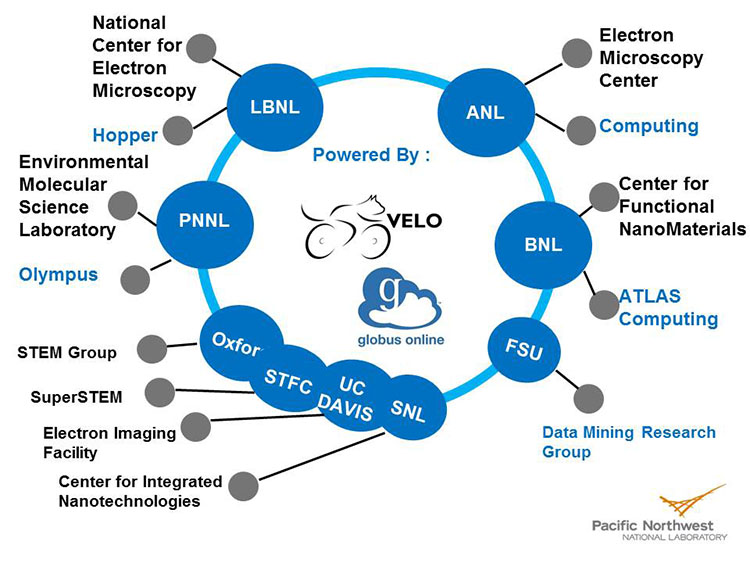
Velo already provides the capability to access registered remote compute and data resources. These capabilities were extended via integration with Globus Share to provide the ability to access new data resources on demand. Again, for the demonstration, the existing Premier Network was enlarged to include microscopists from Lawrence Berkeley National Laboratory (LBNL), ANL, Brookhaven National Laboratory (BNL), Sandia National Laboratories (SNL), and a number of international laboratories. All of the collaborators provided experimental data, while LBNL, BNL, and ANL also contributed data storage and computational resources. Users were able to discover, access, and analyze data stored at all DOE laboratory sites seamlessly, avoiding unnecessary data transfers.
HPC-enabled Data Analysis
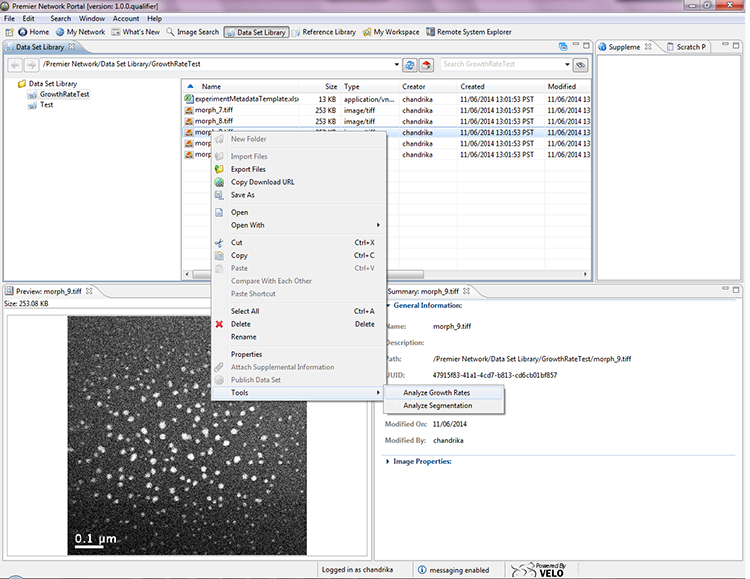
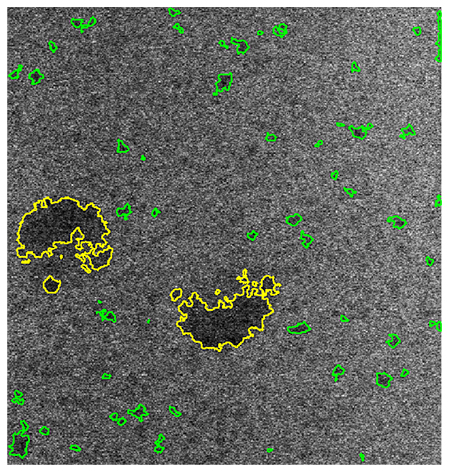
In collaboration with Florida State University (FSU) and the University of Utah (UUtah), two new analysis workflows for single-image and complete video segmentation (including data subsampling) and statistical analysis of particle growth behavior were developed. The Premier Portal associates registered analysis workflows with suitable data set types, offering users only analysis and visualization tools that are appropriate for the data they have. The portal also is aware of which computational facilities currently provide the offered analysis capabilities and will provide users only with the available choices when they initiate their analyses. An easy-to-use screen in the portal provides users with default choices, which will ensure a successful execution of the analysis workflow. Once submitted through the interface, existing Velo capabilities manage job staging, job submission, job monitoring, and returning analysis results to the Premier Portal, which is associated to the original experimental data and with a basic provenance record describing what has been done.
A Demonstration of Scientific Progress
In the demonstration project, an initial study established the experimental viability of coordinating large-scale research efforts at distributed user facilities. After selecting a set of suitable materials using the Materials Project simulation capabilities at the National Energy Research Scientific Computing Center (NERSC), the first test will provide the calibration of the experimental imaging facilities. Each site (LBNL, PNNL, ANL, BNL, and SNL) will provide experimental results for the same nanoparticles from solution and apply the same advanced analysis workflow with software provided by a range of collaborators to establish the nucleation and growth pattern. These results act as a calibration for the facilities and show how experiments performed at different locations and under varied experimental conditions can be used collectively to increase the precision of a fundamental materials experiment. The initial analyses will be extended to a set of new compounds proposed by the Materials Genome project. Using the same tracking algorithms as those used to standardize observations from each facility, trends between the syntheses of different compounds will be identified, e.g., does the overall morphology depend primarily on the precursor, solvent, growth rate, or some combination of these effects, and how do these tie in with the predictions of properties from the genome project?
Notably, the joint data and domain science demonstration development is expected to lead to a number of high-profile publications, which will elevate the impact and visibility of this new type of data science infrastructure provision.
Demonstration Technical Details
For the SC14 demonstrations, the partners demonstrated the following capabilities:
- Remote storage at all sites accessible via Globus Share
- Collaborative data and analysis framework via Velo
- Data access and movement across all sites via Globus Online
- Access to HPC resources at NERSC, PNNL, and ANL
- Access to computational modeling, image simulation, and analysis algorithms from various laboratory, university, and international partners
- Data Publication Service.
An upload of experimental data from the different associated microscopy facilities, by either uploading the data to PNNL or linking to it at local sites using Globus share in connection with Velo, was shown in real time. After the upload, a detailed analysis of the particle growth pattern in the sample run at a number of remote sites was performed. The results of both analytical processes were uploaded and linked to their original data. Finally, by replicating the data to a preferred publication site (e.g., home organization) and requesting a digital object identifier, or DOI, for said data through a publication service, the complete data set was published. If available, these published data were linked to any existing publication.
Progress Since SC14
The new capabilities demonstrated at SC14 have been included in the operational Premier Portal services. Both the easily accessible analytical and data publication capabilities have been of considerable interest to the collaboration. Currently, the teams are working on integrating further analytical capabilities into the portal to offer a broader range of services. SC14 fostered a number of additional collaborations with universities interested in developing analytical tools, and the Premier Portal provides an ideal environment as it affords easy access to a range of scientific data, as well as access to immediate feedback from expert users regarding the tools’ functionality. In turn, Velo offers experts the ability to easily try new tools alongside their existing solution via easy-to-use interfaces (no installation required as all tools are pre-installed server side or on the associated HPC systems).
The Premier Portal has entered into an initial agreement with Springer to coordinate the publication process of one of their journals, Advanced Structural and Chemical Imaging, with the data publication process available through the portal. Scientists can submit extended supplemental information to the portal and publish it with their papers. Once published, others not only can access the data, but they can use the portal-provided analysis and visualization tools to explore the data in situ.
Members of the SC Demonstrator Project
MICROSCOPY: Nigel Browning (PNNL), Andrew Minor (LBNL), Peter Ercius (LBNL), Erich Stach (BNL), Dean Miller (ANL), Katherine Jungjohann (SNL), Hao Yang (Oxford University), Patricia Abellan (SuperSTEM), Wen Tong (University of California, Davis)
COMPUTER SCIENCE: Kerstin Kleese van Dam (PNNL), Michael Ernst (BNL), Hironori Ito (BNL), David Skinner (LBNL), Shane Cannon (LBNL), Rachana Ananthakrishnan (ANL), Ian Foster (ANL), Chiwoo Park (FSU), Liz Jurrus (UUtah), Carina Lansing (PNNL), Bibi Raju (PNNL), Mathew Thomas (PNNL), Chandrika Sivarakrishnan (PNNL), Malachi Schramm (PNNL)
Contact SC14 demonstrators:
- Nigel Browning (Microscopy)
- Kerstin Kleese van Dam (Computer Science)
Premier Portal contact: Terri Clark (User Support)
Acknowledgement
The PREMIER Portal and project is part of the Chemical Imaging Initiative at PNNL. The research was conducted under the Laboratory Directed Research and Development Program at PNNL, a multiprogram national laboratory operated by Battelle for the DOE.