Advanced Computing, Mathematics and Data
Research Highlights
April 2010
PFLOTRAN: Advanced Computing Modeling for a Cleaner Environment
Innovative Computer Code Simulates Movement of Underground Contaminants at Hanford
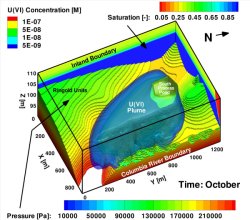
Simulated uranium [U(VI)] plume at the Hanford 300 Area in the October 2009 time frame. Enlarge Image
What are the results? Recent findings employing the PFLOTRAN (petascale reactive multiphase flow and multicomponent transport) code are strengthening scientists' ability to more accurately predict groundwater contaminant movement at the Department of Energy's Hanford site, a former nuclear weapons production complex.
PFLOTRAN has been used to simulate the migration of contaminants in groundwater at the Hanford 300 Area and estimate the release of uranium to the neighboring Columbia River. The model incorporates hydrologic and geochemical reaction parameters derived from field and laboratory experiments to provide an increasingly realistic representation of subsurface physical and chemical processes at the 300 Area site.
For a conceptual model depicting current conditions near the South Processing Pond—where a uranium plume intersects the Columbia and continuous source regions exist—PFLOTRAN estimated that uranium leaches into the river at a rate of 25 kg/year. The estimate is well within the range of estimates based on field studies (i.e., 20-50 kg/year by Peterson et al., 2009).
Stochastic simulations using PFLOTRAN demonstrated that the amount of uranium transported into the river is less sensitive to small-scale (e.g., meter-scale) geologic heterogeneity than originally thought. This result is surprising considering that the geology of the site is composed of heterogeneous cobbles, gravels, and fine sands. This lower sensitivity is likely due to the accumulation of leached uranium over a kilometer-scale region at the river's edge, which is much larger than the scale of the heterogeneity in the model.
Why it matters? The persistence of a uranium contaminant plume in the groundwater at the 300 Area is of concern because concentrations are above EPA defined maximum contaminant levels, and the uranium is slowly leaching into the Columbia River at much lower concentrations. Modeling with PFLOTRAN is providing input to help inform and guide future cleanup decisions.
What are the methods used to achieve the results? Developed as part of a SciDAC-2 groundwater project, PFLOTRAN is written in object-oriented Fortran9X. The finite volume code is based on well-supported frameworks for high-performance computing (i.e., PETSc, HDF5, MPI) and employs domain decomposition to distribute the problem domain across thousands of processor cores. PFLOTRAN is designed to run on computing architectures ranging from conventional laptops and desktops to the world's fastest supercomputers. The code has been run on problems composed of up to two billion unknowns using up to 131,072 processor cores on Argonne National Laboratory's Blue Gene/P and Oak Ridge National Laboratory's Jaguar supercomputers. These resources were made available through the U.S. DOE Office of Science INCITE program.
What are the next steps? Future simulations of the Hanford 300 Area will incorporate increasingly realistic depictions of the Columbia River hyporheic zone—the mud/silt layer along the river bottom. Recent PFLOTRAN simulations have demonstrated that the uranium flux to the Columbia River is highly dependent on the model's representation of the hyporheic zone. The model will incorporate data sets from ongoing field research that better characterize the zone.
Research Team: Glenn Hammond (PNNL) and Peter Lichtner (Los Alamos National Laboratory)
References: Lichtner, P.C. and G.E. Hammond (2010) Placing the Hanford 300 Area IFRC Site in Perspective: Plume Scale Modeling of Uranium Attenuation and Its Flux to the Columbia River, 5th Annual DOE SBR PI Meeting, March 29-31, 2010, Washington DC.
Peterson, R.E., C.F. Brown, and R.J. Serne (2009) Mass Balance Aspects of Persistent Uranium Contamination in the Subsurface at the Hanford Site, Washington, PNNL-SA-67979, October 2009, Pacific Northwest National Laboratory, Richland, WA.
Acknowledgments: This research is supported under the U.S. DOE SciDAC-2 program with funding provided by DOE Offices of Biological & Environmental Research (BER) and Advanced Scientific Computing Research (ASCR). Supercomputing resources were provided by the DOE Office of Science Innovative and Novel Computational Impact on Theory and Experiment (INCITE) program with allocations on NCCS Jaguar at Oak Ridge National Laboratory.
